Speeding up drug discovery with advanced machine learning
Author: Gavin Edwards, Machine Learning Engineer, AstraZeneca
Whatever our job title happens to be at AstraZeneca, we’re seekers. I’m part of the Biological Insights Knowledge Graph (BIKG) team. We help scientists comb through massive amounts of data in our quest to find the information we need to help us deliver life-changing medicines.
AstraZeneca is a research-based biopharmaceutical company headquartered in Cambridge, UK, with strategic research and development (R&D) centers in Sweden, the United Kingdom and the United States. The company has a broad portfolio of prescription medicines, primarily for the treatment of diseases in three therapy areas — Oncology; Cardiovascular, Renal & Metabolism; and Respiratory & Immunology.
Finding new ways to speed the discovery and development of new medicines
At AstraZeneca, our drug discovery and development is guided by our ‘5R Framework’: right target, right patient, right tissue, right safety, right commercial potential. Over a five-year period, this approach has enabled us to achieve a more than four-fold improvement in the number of pipeline molecules advancing from pre-clinical investigation to completion of Phase III clinical trials.
At the same time, the vast amount of data our research scientists have access to is exponentially growing each year, and maintaining a comprehensive knowledge of all this information is increasingly challenging.
Machine learning is the key to analyzing data quickly and finding relevant connections. That’s why our team is using knowledge graphs, which are networks of contextualized scientific data facts such as genes, proteins, diseases, and compounds — and the relationship between them — to give scientists new insights.
How the AstraZeneca Biological Insights Knowledge Graph (BIKG) group uses PyTorch and Microsoft Azure Machine Learning
Our team is a mix of Graph Engineers, Data Scientists, Bioinformaticians, and Machine Learning Engineers. We are all working together to build a data pipeline that assembles scientific data into a knowledge graph, which is then used to train machine learning models for early drug discovery decisions at AstraZeneca. We’ve combined research in the public domain and our internal research into a graph that encodes complex information easily. By layering machine learning on top of that, we can train machine learning models that recommend novel drug targets and help to inform pipeline decisions. We can use this approach to identify, say, the top 10 drug targets our scientists should pursue for a given disease.
Because a great deal of the data is unstructured text, we need a way to extract relevant data from it. That’s where our Natural Language Processing (NLP) team comes in. They use a variety of neural network architectures and rule-based methods for biomedical text mining. The NLP team members like to use PyTorch, an open-source machine learning library developed from the Torch library, to define and train models based on our needs. PyTorch is a natural choice for teams that want to be at the forefront of NLP research, as it’s widely used by the academic community and allows us to quickly implement ideas from the latest papers. By using PyTorch in our projects, we have been able to quickly adopt the latest research and build customized NLP models. We look forward to integrating and improving our projects with PyTorch.
We also use PyTorch with Azure Machine Learning for our recommendation systems, where we train embeddings and machine learning models on our knowledge graph. Our scientists use embeddings to map nodes in the graph, such as a drug or disease, into a low-dimensional numerical form that meaningfully represents the original data. We can then leverage these embeddings to make recommendations and train use case–specific models. Many alternative technologies for training machine learning models require that you set them up yourself, or they don’t offer the built-in support for the open source tools and framework that we get with Azure Machine Learning.
Azure Machine Learning has compute capabilities that help us scale training as well as resource management. The PyTorch community then provides us with the various tools for fundamental research and training models. This allows us to build quickly and focus on the science. For example, Azure Machine Learning and PyTorch were used in our NLP team’s recent paper on comparing how our models perform under different circumstances (https://f1000research.com/articles/9-710). We need to think operationally when dealing with complex processes and huge amounts of compute. We use Azure Blob storage to handle the many terabytes (currently about 5 TB and growing) of data used and created by our experiments. We scale our Azure Machine Learning use to minimize the cost of our compute. When we’re not training, the compute auto scales down to zero, saving us cost. And we also can use that elasticity to make sure that we don’t have devices in queue, waiting to use compute time, so it’s faster. When we have a very demanding compute job, we can quickly scale up to a large number of compute instances.
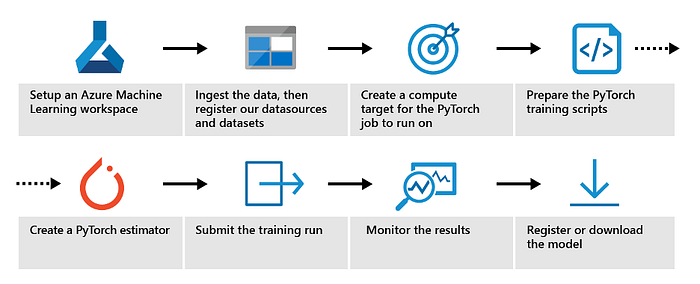
Another benefit is the end-to-end ML lifecycle management, such as dataset and model management. We use these features to efficiently organize our library of models and deploy them against different data, tracking their performance via the interface. This speeds up our iterations and ultimately leads to faster model development. Successfully creating machine learning models involves many steps, from the initial decision for model architecture through training and tuning.
Finally…
AstraZeneca applies the models we build using this approach to try and find potential new and novel drug targets faster and to make better recommendations. We’re focused on saving time for scientists and delivering other advantages, too. We want to enable our scientists to explore the knowledge graph when they want to test a certain theory, confirming or excluding that theory quickly within the literature, and then investigating further.
Just as we typically shrink a knowledge graph down to explore one dimension of a problem, we can expand it to give scientists a wider view to break through silos. For example, an expert on lung cancer might benefit from using the knowledge graph to expand their view into other areas, gathering new insights and information that could be valuable to their research. The flexibility and reach we’re now able to give them can inspire more creativity.
So although we combined these two technologies to quickly and cost-effectively target early drug discovery decisions and recommendations, the scope is much bigger than that. There’s so much more you can do.
What’s next
We continue to expand our knowledge graph. And we see so many opportunities to apply machine learning in the pharmaceutical industry where it can take 10 to 15 years to go from an idea for a drug to a medicine that’s ready to go to market. If we can increase the speed and efficiency of that process, we can hopefully deliver new medicines to the patients and healthcare professionals who rely on us.
About the author

I joined AstraZeneca last year as a Machine Learning Engineer within the BIKG team. Our team is a group of talented people with a variety of skill sets. My academic background is in Computer Science with industry experience in Data Science and Cyber Security. I am currently developing the machine learning models that exploit our knowledge graph.
We have many exciting opportunities available at AstraZeneca, and we are always on the lookout for talented people. If you are interested in joining us, why not take a look at our current vacancies?


